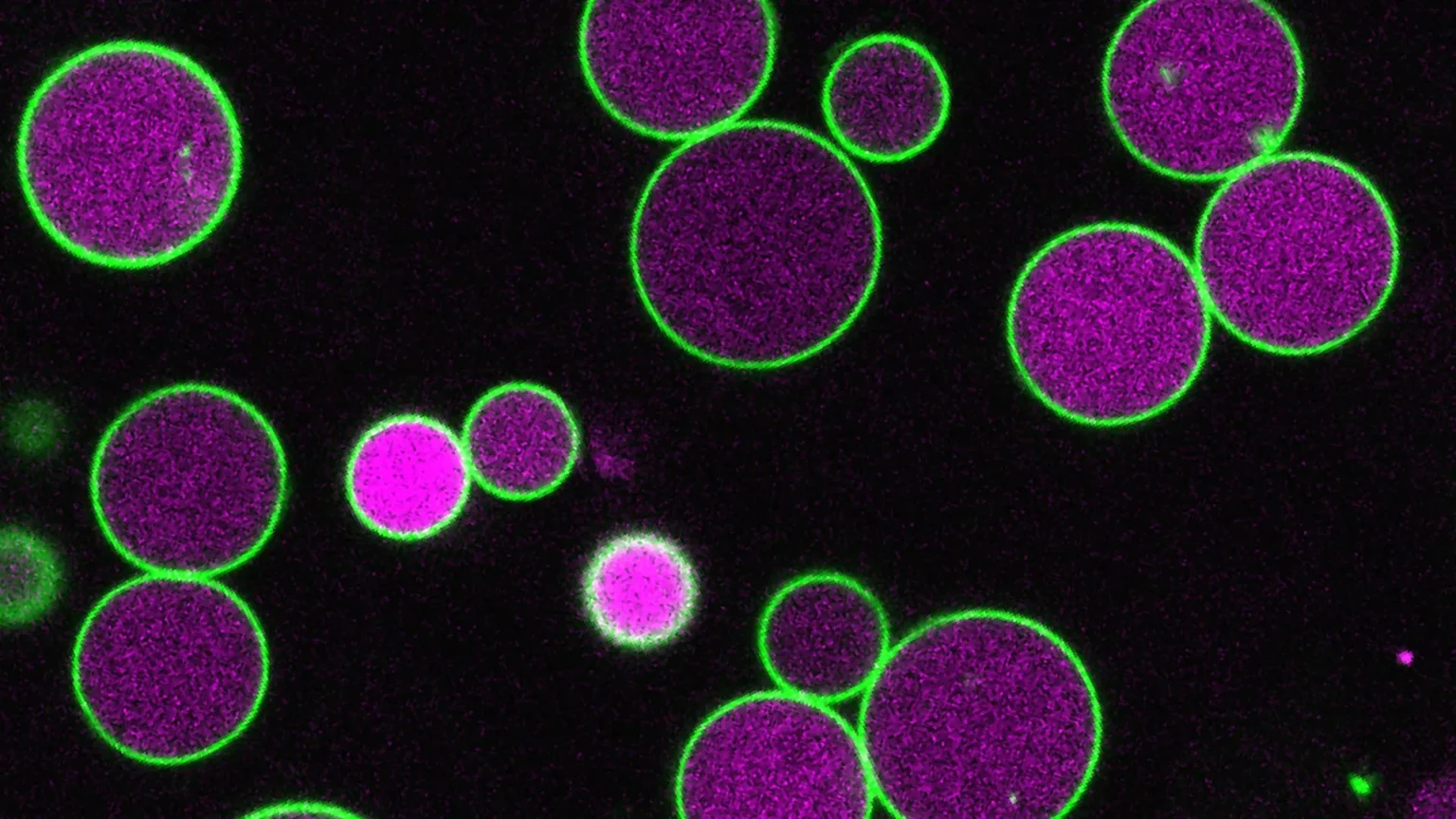
Plants spend most of their lives using photosynthesis to create energy. However, in the early stages after the seeds begin to grow, they are not yet able to capture light. During this short but important period, they instead rely on stored fatty acids. To break down these fatty acids, plant cells use specialized structures known as peroxisomes. Peroxisomes are membrane-bound compartments that are also found in human cells. Due to their size and visibility, plant cells serve as a useful system to study how peroxisomes function.
“The plant we use, Arabidopsis, has large cells and peroxisomes that you can see inside with a light microscope,” said Bonnie Bartel, Ralph and Dorothy Rooney Professor of Biological Sciences. “Peroxisomes grow larger during the seed-to-seedling stage, when plants rely on fatty acids for energy, and then shrink to their normal size once the plant is able to photosynthesize.”
Protein PEX11 helps control peroxisome size
Bartel’s team focuses on these enlarged peroxisomes, specifically a protein called PEX11. Scientists have long known that PEX11 plays a role in helping peroxisomes divide. In a new study published in Nature Communications, the researchers found that this protein also helps control the expansion and contraction of peroxisomes during early plant development.
“Peroxisomes have been implicated in some human diseases and are used in bioengineering,” said lead author Nathan Tharp, a Rice graduate student. “But they can be quite difficult to study.”
Study complex proteins using CRISPR
A common strategy for understanding proteins is to disable the genes involved in protein production and observe the effects. In this case, the situation was even more complicated. PEX11 is produced by five different genes. Destroying just one of them will have little effect, but removing all five will cause the plant to die. This has made it difficult to accurately identify the protein’s function.
To get around this problem, Tharp used advanced CRISPR technology to selectively disable different combinations of five genes.
“Using recent advances in CRISPR, we were able to disrupt a specific combination of five genes,” said Tharp, who recently defended his paper. “For the first time, we found that PEX11 is clearly involved in controlling peroxisome growth from the seed to seedling stage.”
Giant peroxisomes reveal growth control mechanisms
Tharp designed two types of mutant plants, each lacking a specific set of PEX11 genes. In both cases, peroxisomes expanded from the seed to seedling stage, as expected. However, some individuals never returned to normal size and continued to grow far beyond their normal limits. In extreme cases, peroxisomes extended from one end of the cell to the other.
These mutant cells also lacked vesicles, small membrane-bound compartments that normally form inside peroxisomes during fatty acid treatment. Under normal conditions, these vesicles develop as the peroxisome grows and appears to remove part of its outer membrane.
“Vesicles that engulf part of the membrane as they form may help control peroxisome growth,” Tharp said. “In our PEX11 mutants, these vesicles either don’t form or are abnormally small and rare, so we see these giant peroxisomes that are much larger than normal.”
Discoveries extend beyond plants to other species
Although the research focused on plants, Tharp wanted to know whether the same mechanism existed in other organisms. To test this idea, he introduced a yeast version of a protein called Pex11 into mutant plant cells.
“We put yeast Pex11 into mutant plant cells to see if we could restore peroxisomes to normal,” Tharp said. “And so it happened.”
This result suggests that Pex11 functions similarly in yeast as in plants, despite the long evolutionary distance between the two. Therefore, this protein may play an equivalent role in other cell types, including human cells.
“The fact that this protein was found to play the same role in yeast and plant cells suggests that it may be a highly conserved protein,” Bartel said. “Therefore, our findings in plants in this relatively easy-to-study model may also be applicable to human cells and cells used in bioengineering.”