Optibrium, a leading developer of software and AI solutions for molecular design, today announced the new QuanSA™ plugin for PyMOL. It provides an intuitive graphical user interface (GUI) for the ligand-based binding affinity prediction methods that are part of the company’s BioPharmics 3D molecular modeling platform. The new interface gives chemists easier access to accurate affinity predictions to guide the design of potent compounds, reducing the synthesis and testing burden of lead optimization.
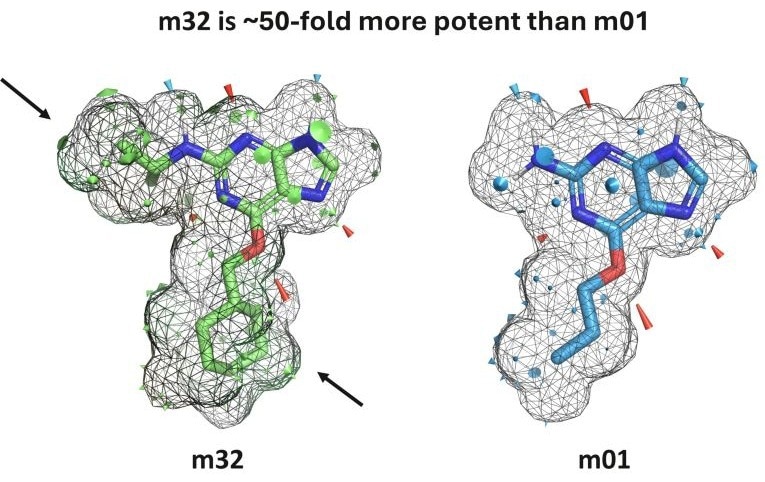
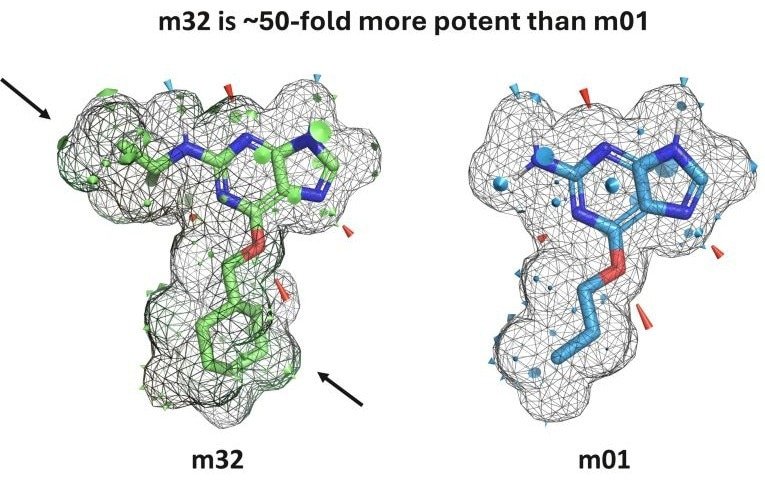
An example of the visual output provided by the new QuanSA PyMOL plugin. m32 is approximately 50 times stronger than the structurally similar m01, despite having an almost identical hydrogen bonding pattern (red and blue cones). This difference is explained by the additional steric contribution indicated by the surface patch highlighted by the black arrow. Image credit: Optibrium Ltd.
QuanSA (Quantitative Surface Field Analysis) was originally developed as a command line tool for computational professional users, but is now available to the broader chemical community as a new PyMOL plugin. The plugin’s clear visualization identifies key interactions that drive molecule affinity, providing key insights that allow users to optimize molecule potency.
QuanSA is a differentiated and validated method for predicting the affinity of potential drug molecules for biological targets. Its physically motivated machine learning approach explicitly models the factors governing molecular recognition and binding. This provides accuracy comparable to leading simulation-based methods such as free energy perturbation (FEP), but at a fraction of the computational cost and without the need for protein structure. QuanSA makes accurate affinity predictions available much earlier in a project and allows these predictions to be applied to a larger number of compounds and a wider range of targets.
The QuanSA plugin follows the recent introduction of a PyMOL interface for Surflex-Dock, Optibrium’s molecular docking method, and reflects the company’s continued efforts to make advanced 3D modeling techniques more accessible. The command-line interface remains fully supported for professional users and large-scale screening applications.
Early stage drug discovery relies on accurate prediction of binding affinity. QuanSA has been shown to provide accuracy comparable to state-of-the-art simulation-based methods, but at a fraction of the computational cost and can be achieved even when protein structures are not available. Providing this functionality to the broader scientific community through an intuitive and visual interface is an important step. The more broadly applicable these predictions are, the greater their impact on drug discovery. ”
Ann Cleves, Vice President, Application Science, Biopharmaceuticals, Optibrium
Matthew Segal, CEO of Optibrium, added: “Understanding not just how tightly a molecule binds, but also why it binds to a target, is extremely valuable in lead optimization. Using the new PyMOL plugin, the team can now use QuanSA The ability to visualize the key interactions driving affinity alongside proven predictions provides insight to make better and more confident design decisions, resulting in a more informed and efficient path to preclinical candidates.”
The QuanSA plugin for PyMOL is available to BioPharmics license holders at no additional cost.