More than 64,000 proteomics datasets are currently flowing through ProteomeXchange, and the consortium’s latest updates demonstrate how smarter standards, powerful reuse tools, and AI-enabled resources are reshaping biological data sharing.
 Database Update: ProteomeXchange Consortium 2026: Making Proteomics Data Fair. Image credit: Christoph Burgstedt /Shutterstock
Database Update: ProteomeXchange Consortium 2026: Making Proteomics Data Fair. Image credit: Christoph Burgstedt /Shutterstock
In a recent database update paper published in a journal Nucleic acid researchan international team of authors discussed recent advances, data growth, standardization, and future directions of the ProteomeXchange consortium, which enables FAIR (Findable, Accessible, Interoperable, Reusable) proteomics data sharing.
Background of proteomics data sharing and FAIR principles
What happens when thousands of biological datasets go unused? In proteomics, data sharing is essential to advancing disease, drug, and human biology research. Over the past decade, the rapid rise of mass spectrometry-based proteomics has generated vast datasets, whose value depends on accessibility and reuse. The FAIR principles were developed to guide the management and management of scientific data in a way that supports reproducible and transparent science. Collaboration platforms now play a key role in integrating and distributing such data across disciplines. However, continuous innovation is required to address the increasing complexity of new datasets.
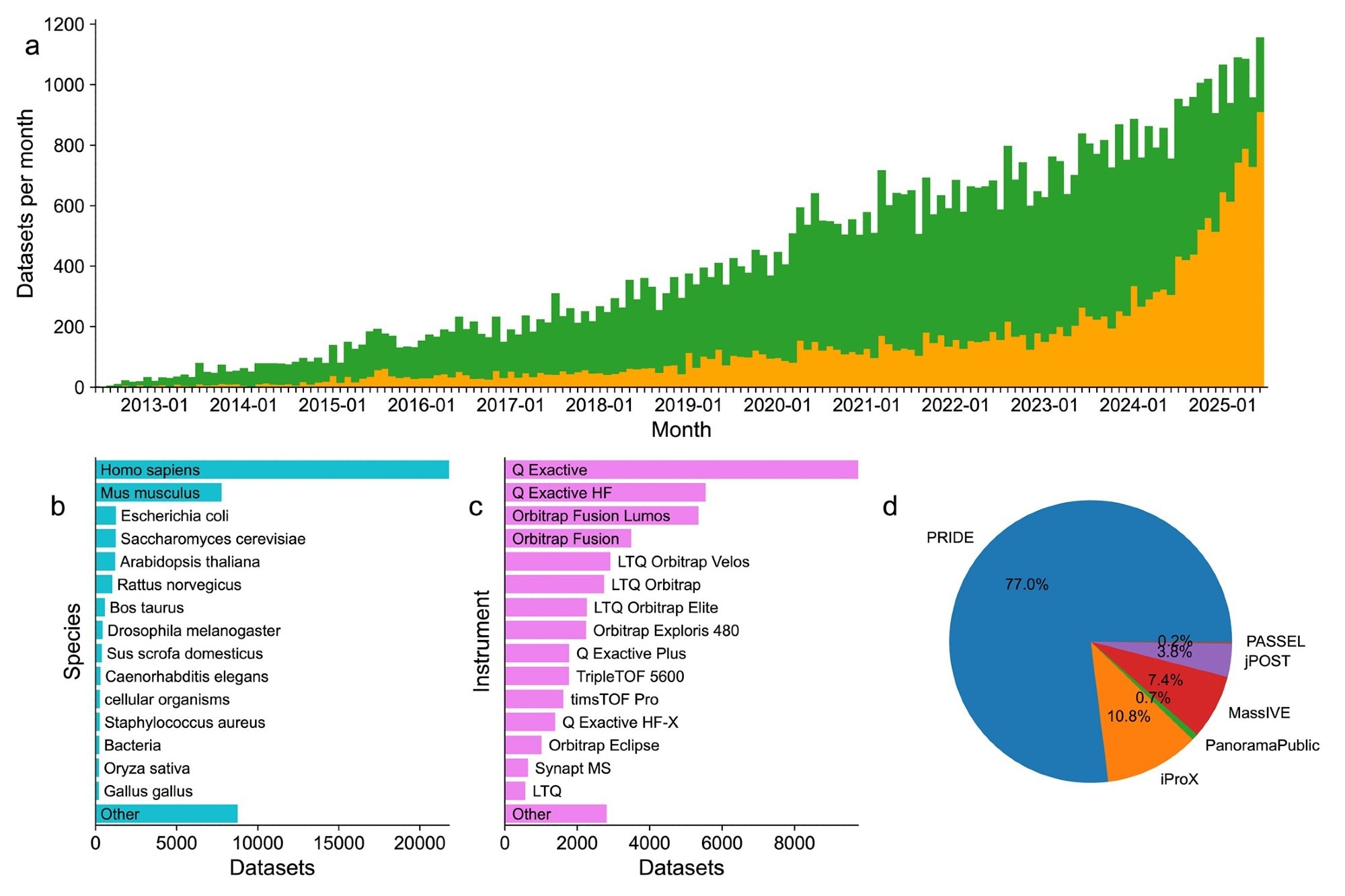
Summary statistics for datasets deposited to the ProteomeXchange resource since 2012. (A) Trends in published (green) and unpublished (orange) datasets from May 2012 to June 2025. A total of 1156 datasets were submitted in June 2025. (B) Summary of the top 15 species in datasets published since 2012. (C) Summary of the top 15 species reported by submitters of datasets published since 2012. (D) Summary of relative counts of all datasets by receiving repositories.
ProteomeXchange Infrastructure and Data Standards
This consortium maintains an infrastructure that enables standardized submission, storage, and dissemination of proteomics data generated by mass spectrometry. Member repositories that contributed to data archiving and access include the PROteomics IDentifications Database (PRIDE), PeptideAtlas, Mass Spectrometry Interactive Virtual Environment (MassIVE), Japan Proteome Standards Repository/Database (jPOST), Integrated Proteome Resource (iProX), and Panorama Public. The submitted dataset consisted of raw mass spectrometry files, processed data including identification and quantification results, and experiment metadata structured according to standards developed by the Proteomics Standards Initiative (PSI).
Efficient uploads were performed using a number of data transfer protocols, including File Transfer Protocol (FTP), Aspera, Hypertext Transfer Protocol Secure (HTTPS), Web Distributed Authoring and Versioning (WebDAV), and PRESTO. Additionally, metadata standardization has been improved through the Sample and Data Relationship Format (SDRF)-Proteomics, allowing for clear mapping between samples and experimental conditions. A unique dataset identifier (ProteomeXchange dataset identifier) ensured traceability, and reanalyzed datasets were assigned an RPXD identifier.
ProteomeCentral integrates metadata from all repositories and enables search and retrieval of datasets through a single platform. Universal spectral identifiers (USIs) have enabled precise identification and visualization of single spectra. This infrastructure also facilitates large-scale reuse, integration with external resources, and use in machine learning and artificial intelligence (AI) workflows.
ProteomeXchange Growth, Reuse, and AI Applications
The latest submission statistics from the consortium showed a significant increase in the sharing and reuse of proteomics data worldwide. By June 2025, a total of 64,330 datasets had been submitted, of which 44,248 (69%) were publicly available, reflecting a strong commitment to open science. Notably, 47% of all datasets were submitted within the past three years, highlighting an accelerating trend in data generation and sharing.
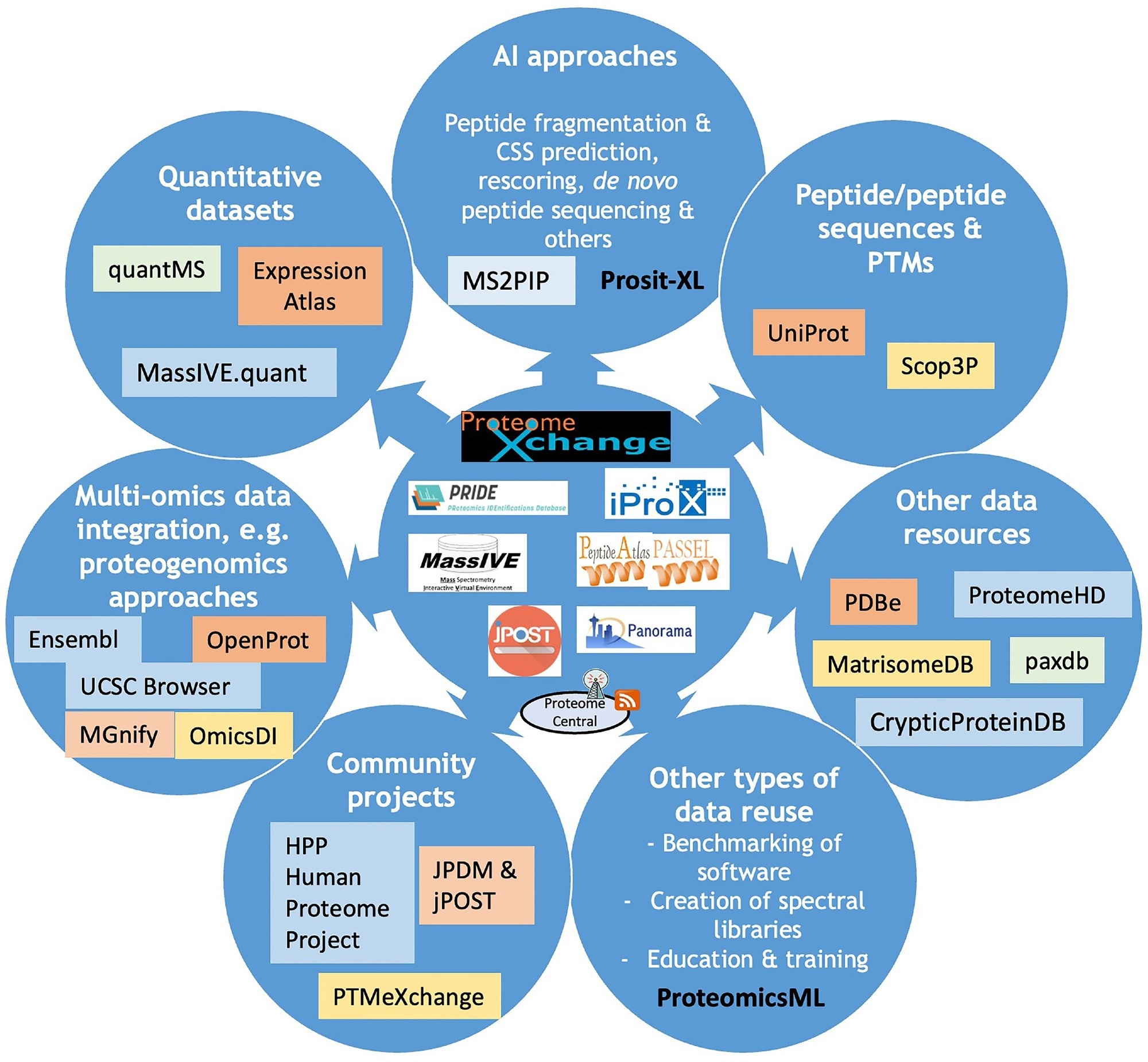
Overview diagram including current ProteomeXchange resources and key efforts spent on data reuse of public proteomics datasets. Different types of data reuse are listed, and for each the corresponding tools and data resources that can access these data are indicated.
Most posts came from the PRIDE repository (77%), followed by iProX (11%), MassIVE (7.4%), jPOST (3.8%), and small amounts from Panorama Public and PeptideAtlas. More than 80 countries have contributed to these public proteomics resources, demonstrating the widespread use of proteomics in biomedical research worldwide.
ProteomeXchange resources increasingly support standardized formats and richer metadata, enhancing interoperability between datasets. The format developed by PSI and SDRF Proteomics has improved and enhanced the quality, reproducibility, and value of the dataset’s metadata. The overall use of USI has facilitated access and visualization of individual spectra in multiple different data repositories. This provides greater transparency and validation of experimental results.
Data reuse activity also increased across the consortium. Public datasets were reanalyzed to gain new biological insights, including protein sequence validation and identification of post-translational modifications. Integration with the UniProt Knowledge Base (UniProtKB) enabled mapping of over 93% of the human proteome, demonstrating the power of data analysis.
Quantitative proteomics resources such as MassIVE.quant and quantms have enabled reproducible large-scale analysis. Additionally, multi-omics integration through resources such as Omics Discovery Index (OmicsDI) and MGnify has enabled the integration of proteomics, genomics, and transcriptomics datasets.
Artificial intelligence and machine learning applications are increasingly supported by the availability of high-quality datasets. Tools such as MassIVE-Knowledge-Base (MassIVE-KB) and ProteomicsML have enabled the development of predictive models for peptide identification, fragmentation, and protein quantification. These advances are turning proteomics into a data-driven field with potential for future applications in precision medicine.
Many challenges still exist in this research area. Due to privacy regulations such as the General Data Protection Regulation (GDPR) and the Health Insurance Portability and Accountability Act (HIPAA), human data requires more access control systems and repository capabilities. Additionally, new techniques have emerged that use proteomics as the primary measurement method and are mass spectrometry-independent, including affinity proteomics platforms such as SomaLogic and Olink assays. This will lead to new research methodologies. Therefore, researchers may require additional resources.
Future directions for the FAIR proteomics infrastructure
The ProteomeXchange consortium has created an innovative and collaborative environment for sharing proteomics data globally, in line with FAIR principles. The introduction of standardized formats, increased scalability, and provision of cutting-edge analytical tools have facilitated widespread reuse of existing data and fostered innovation in biology and medicine. However, future progress will depend on solving data privacy, scalability, and new technologies.
There is a continued need for innovation and collaboration to maintain broad accessibility and support the continued reliability and impact of proteomics data in advancing scientific discovery and enabling broader bioinformatics reuse.
sauce:
Reference magazines:
- Deutsch, E. W., Bandeira, N., Perez-Liberol, Y., Sharma, V., Carver, J. J., Mendoza, L., Kundu, DJ, Bandura, C., Kamatchinathan, S., Hewa. Pathirana, S., Sun, Z., Kawano, S., Okuda, S., Connolly, B., McLean, B., Makkos, M.J., Chen, T., Zhu, Y., Ishihama, Y., and Vizcaino, J.A. (2026). ProteomeXchange Consortium 2026: Fairing Proteomics Data. Nucleic acid research. 54(D1). D459 to D469. Doi: 10.1093/nar/gkaf1146, https://academic.oup.com/nar/article/54/D1/D459/8315797