Scientists reveal how evolution within our own tissues causes disease, protects cells and reveals hidden therapeutic targets for future precision medicine.
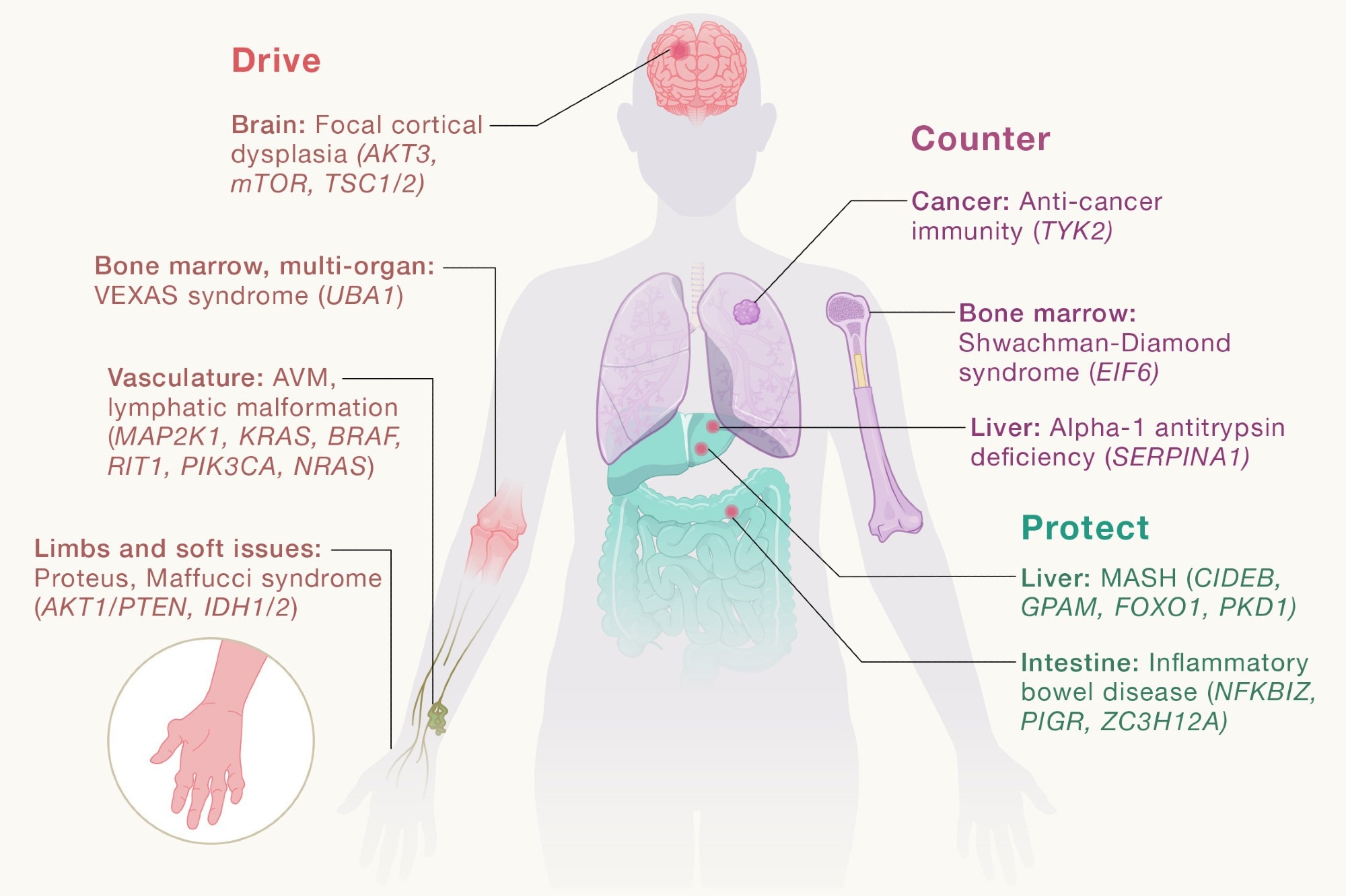
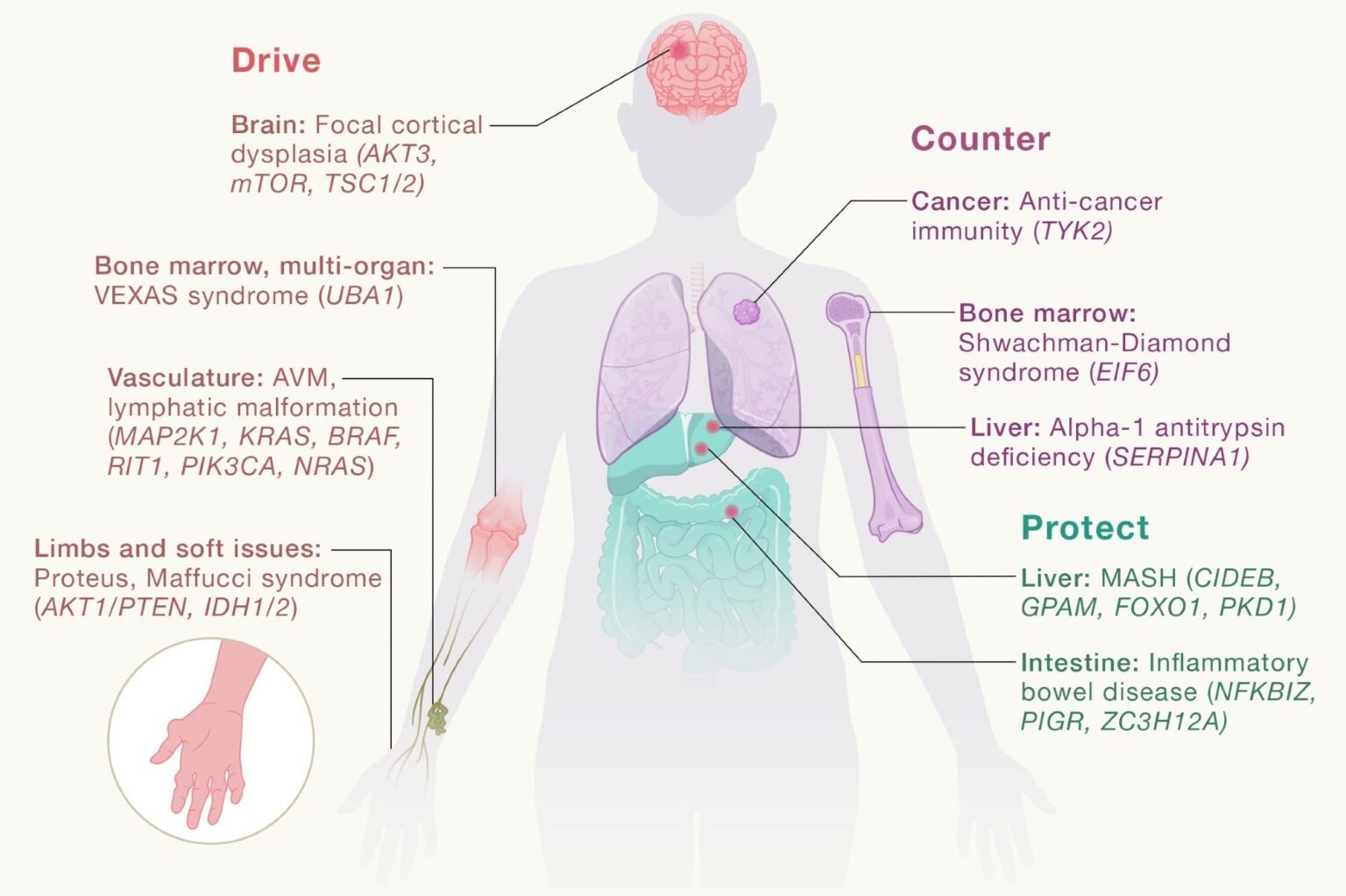
Somatic cell genomics reveals the consequences of evolutionary competition within tissues that can drive disease, counter monogenic diseases, and protect against common diseases.
In a recent study published in the journal cellresearchers reviewed the current evidence on somatic mutations in disease and its potential applications in biomedical discovery.
Somatic mutations, genetic changes that occur in cells after conception, are widespread in healthy tissues. Although most mutations are functionally unimportant and do not change the cell’s phenotype, some mutations occur in important regulatory elements or coding regions within the genome and affect the phenotype. Darwinian selection acts on such phenotypic variation and can lead to the expansion or disappearance of clones with altered fitness.
As a result, normal tissues may display clones with actively selected somatic mutations, called driver mutations. These driver mutations overlap between malignant and normal tissues in many organs, indicating that the earliest events of cancer can be detected in normal tissues. Nevertheless, many driver mutations in normal tissues are absent or infrequent in cancer and do not necessarily contribute to malignant transformation.
Disease processes constitute additional selective pressures on mutant clones. Research suggests that disease often selects for a different set of driver mutations than those observed in cancer or healthy tissue, and that clones grow to larger sizes in diseased tissue than in healthy tissue. In this study, the researchers discuss how somatic mutations shape disease biology, sometimes contributing to pathology but other times conferring an adaptive advantage to affected cells, while highlighting that mutations that benefit individual clones do not always improve tissue- or organism-level health, and outline potential implications for clinical applications.
Factors shaping somatic cell mosaicism and clonal expansion
Organ-specific structures may determine the extent of somatic cell diversity. The hematopoietic system has few spatial and geographic limitations. Blood variants that yield the highest fitness effects can be scaled up without space constraints. In contrast, hepatocytes are restricted by lobular borders, which may act as a structural constraint to clonal expansion and may represent a fibrotic barrier in chronic liver disease.
Furthermore, inflammation is one of the selective pressures that can increase constant potential clonal hematopoiesis (CHIP). Inflammation has been shown to promote clonal expansion of tet methylcytosine dioxygenase 2 (TET2) mutant cells in rodents through tumor necrosis factor (TNF)-α and interleukin (IL)-6 activity. Some chemicals act as carcinogens by increasing mutagenesis.
Carcinogens can also select clones through promoter mechanisms. Many carcinogens, which are considered mutagens, promote cancer by altering the selective landscape of tissues and promoting clonal proliferation. Contamination promotes lung cancer development by increasing inflammation through IL-1β and may lead to the expansion of existing mutant Kirsten rat sarcoma virus oncogene homolog (KRAS) clones rather than directly generating their mutations.
Somatic mutations as a cause of human disease
Somatic mutations are emerging as a cause of certain autoimmune and neurological diseases, namely diseases with idiopathic etiology, such as autoimmune diseases and focal epilepsy. Malformations of cortical development often accompany refractory epilepsy and are primarily caused by somatic mutations acquired during development, primarily activating mutations in the phosphoinositide 3-kinase (PI3K), protein kinase B (AKT), and mechanistic target of rapamycin kinase (mTOR) pathways.
Arteriovenous malformations are vascular abnormalities in which veins and arteries are improperly connected without a normal capillary bed. Often they are caused by somatic mutations in the rat sarcoma, mitogen-activated protein kinase (RAS, MAPK) pathway. Maffucci syndrome and Aurier disease are non-inherited skeletal diseases characterized by hemangiomas and enchondromas, both of which are associated with somatic mutations in isocitrate dehydrogenase 1 (IDH1) or IDH2.
Adaptive somatic mutation and disease prevention
Somatic mutations can fight or protect against disease.
Many adaptive somatic mutations have the potential to counter or alleviate disease-associated cellular stress, providing potential therapeutic opportunities. In inflammatory bowel disease (IBD), recurrent somatic mutations have been identified in genes involved in IL-17 signaling in intestinal tissues. Modeling has revealed that these mutations render intestinal cells resistant to the deleterious effects of IL-17-mediated inflammation at the cellular level, but their ultimate impact on disease progression in patients is still being investigated.
CHIP mutations can worsen the disease or predict the development of leukemia, but in some cases they may also be protective. In the context of bone marrow transplantation, CHIP within the donor bone marrow can benefit recipients by increasing survival and reducing recurrence rates in certain clinical situations. CHIP has also been reported to be associated with improved response to immunotherapy in several cancer types. Additionally, adaptive somatic mutations have been detected in cirrhotic livers.
Somatic mutations in AT-rich interaction domain 1A (ARID1A), polycystin 1 (PKD1), and lysine methyltransferase 2D (KMT2D) may increase cell fitness and protect cells from damage. They can also promote liver regeneration after injuries such as chemical toxins or surgical resection by promoting clonal survival and expansion of affected hepatocytes, although adaptive benefits at the clonal level do not always translate into improved outcomes for the whole organism. Adaptive somatic mutations can arise under selective pressure due to germline mutations that cause monogenic diseases. In such cases, somatic mutations may counter germline mutations and partially restore cellular function in the affected tissue.
Somatic cell genomics as a discovery framework
Somatic cell genomics may represent an alternative and complementary approach to germline genetics. Cancer genomics research is some of the earliest and most extensive somatic cell genomics research, providing a roadmap for non-malignant diseases. Although the landscape of somatic mutations in most diseases and tissues is poorly understood, preliminary studies indicate that somatic mutations may reveal biologically relevant pathways and potential therapeutic targets, especially when analyzing patterns of positive selection among clones to identify genes under evolutionary pressure in diseased tissues.
A proposed framework for somatic cell genomics, driven target discovery
The authors propose a four-step framework for systematic target discovery leveraging somatic cell genomics. This framework includes 1) selection of cells based on phenotypic or cellular markers, 2) sequencing of somatic mutations, 3) deciphering selection patterns to identify candidate genes, and 4) validation of genetic findings to name drug targets.
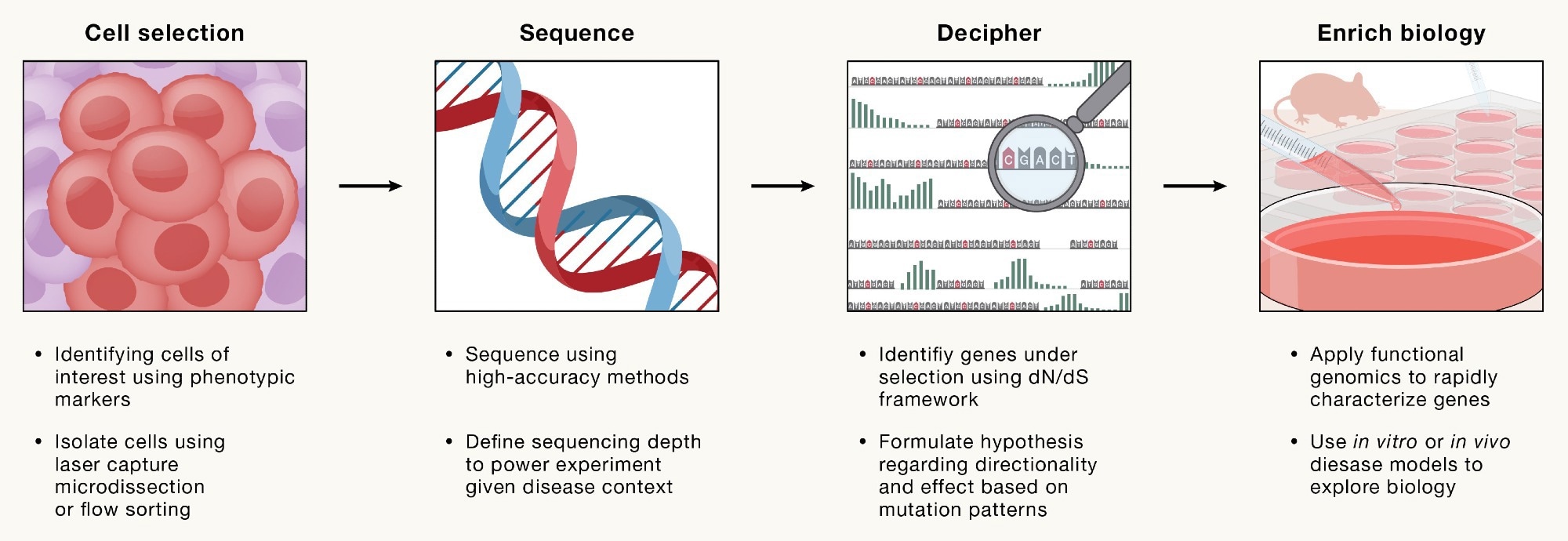
A four-step framework for systematically identifying somatic gene targets that impact disease and inform therapeutic strategies.
Somatic cell genomics offers a promising strategy to elucidate disease mechanisms and identify therapeutic targets, although careful experimental validation and interpretation of effects at the clonal and organismal levels will be required before clinical application.